Colorectal cancer is estimated to cause more than 53,000 deaths in the US in 2020, making it the second most common cause of cancer death. This death rate, however, has been steadily dropping, likely due to better cancer screening programs such as colonoscopy. During colonoscopies, clinicians excise colorectal polyps and visually examine them on histopathology slides for neoplasia. Detecting cancer at an early, curable stage and removing pre-invasive adenomas or serrated lesions ultimately reduces the mortality rate. Numbers and types of polyps found can also indicate future risk for malignancies and are therefore used as the basis for screening recommendations.
An artificial intelligence (AI) model for automated classification of colorectal polyps could benefit cancer screening programs by improving efficiency, reproducibility, and accuracy, as well as reducing access barriers to pathological services. In a new study out of Dartmouth’s and Dartmouth-Hitchcock’s Norris Cotton Cancer Center, a computer science and clinical research team led by Saeed Hassanpour, PhD, trained a deep neural network to do just that. Not only can their model distinguish the four major types of colorectal polyps at the level of practicing pathologists, as evaluated on a dataset across multiple external institutions, but also proves that a model designed using data from a single institution can achieve high accuracy on outside data.
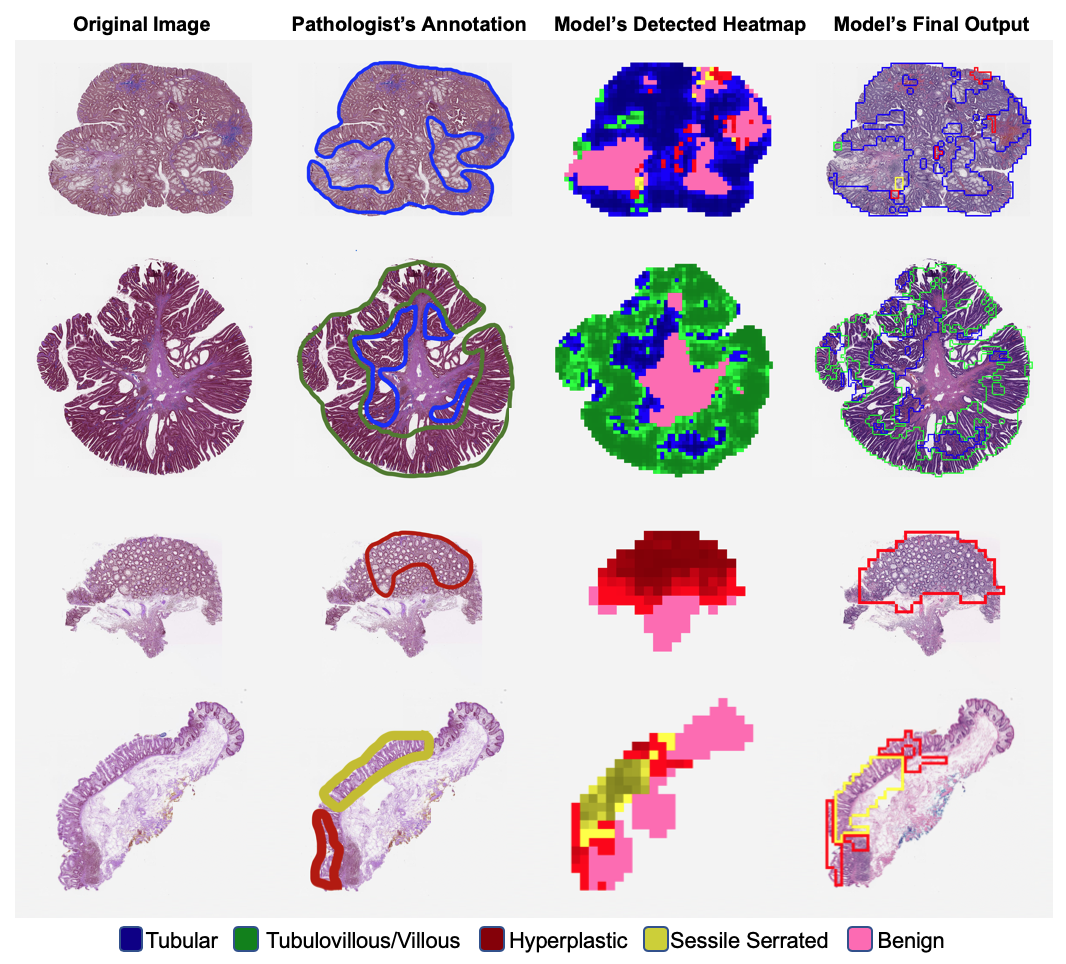 The team found that a deep neural network, trained on colorectal polyp data from Dartmouth-Hitchcock Medical Center, still performed with the same level of sensitivity and accuracy as practicing pathologists when used on 238 slides spanning 24 different institutions in the US. These findings, “Evaluation of a Deep Neural Network for Automated Classification of
The team found that a deep neural network, trained on colorectal polyp data from Dartmouth-Hitchcock Medical Center, still performed with the same level of sensitivity and accuracy as practicing pathologists when used on 238 slides spanning 24 different institutions in the US. These findings, “Evaluation of a Deep Neural Network for Automated Classification of
Colorectal Polyps on Histopathology Slides,” have been published in JAMA Network Open. “Our study is one of the first to show a deep neural network that is generalizable to data from multiple external medical centers,” says Hassanpour. “A challenge in the field of deep learning for medical image analysis is collecting widespread data. Here, we have access to histopathology slides from 24 different institutions, which gave us the opportunity to evaluate and show that the AI models that we train are broadly generalizable to new data from outside.”
The access to a multi-institutional dataset was made possible by Hassanpour’s collaboration with Dr. Arief Suriawinata, MD, and his group from the Department of Pathology & Laboratory Medicine at Dartmouth-Hitchcock Medical Center and Dr. Elizabeth Barry, PhD, from the Department of Epidemiology at the Geisel School of Medicine at Dartmouth, as well as her colleagues from the Vitamin D/Calcium Polyp Prevention Clinical Trial.
Hassanpour’s team has built a graphical user interface for showing the classifications of the neural network. They are currently working on a clinical trial to evaluate the use of their algorithm for assisting pathologists in diagnosis of colorectal polyps. “We are hoping to create a software application that can help pathologists improve their accuracy, efficiency, and consistency in diagnosing slides,” he says.
Saeed Hassanpour, PhD, is an Associate Professor of Biomedical Data Science and Associate Professor of Epidemiology at the Geisel School of Medicine at Dartmouth, Associate Professor of Computer Science at Dartmouth’s College of Arts and Sciences, and member of the Cancer Population Science Research Program at Dartmouth’s and Dartmouth-Hitchcock’s Norris Cotton Cancer Center. His research interests include biomedical informatics, machine learning, and personalized medicine. https://www.hassanpourlab.com/ @SaeedHP