 My research focuses on understanding the mechanism by which motors and structural proteins organize microtubules at spindle poles.To this end, I have written a computer simulation using the physical, chemical and biological properties of these motor and structural proteins. We used previously published biophysical data as parameters for each of the proteins, and trained the simulation so that its output matched the results of immunodepletions performed in an extract system. Using this simulation, we were able to generate a simple mechanistic model for microtubule organization at spindle poles that involves a balance between motor forces and crosslinking activity. The model made certain specific predictions, which were confirmed with experiments performed in a mitotic extract.
My research focuses on understanding the mechanism by which motors and structural proteins organize microtubules at spindle poles.To this end, I have written a computer simulation using the physical, chemical and biological properties of these motor and structural proteins. We used previously published biophysical data as parameters for each of the proteins, and trained the simulation so that its output matched the results of immunodepletions performed in an extract system. Using this simulation, we were able to generate a simple mechanistic model for microtubule organization at spindle poles that involves a balance between motor forces and crosslinking activity. The model made certain specific predictions, which were confirmed with experiments performed in a mitotic extract.
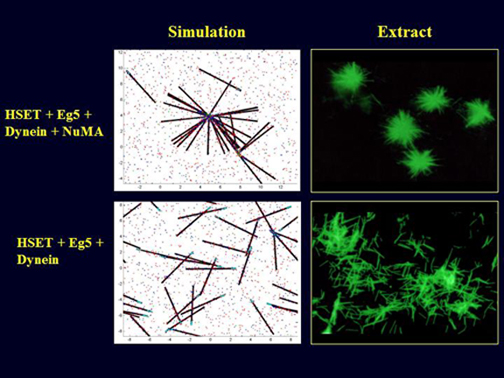 TOP ROW: Comparison of an output of the simulation showing an aster (a radially arranged array of microtubules) and associated motor proteins (left panel) vs asters formed in a mitotic extract under the same conditions (right panel, immunofluorescence staining is for tubulin, showing microtubules)
TOP ROW: Comparison of an output of the simulation showing an aster (a radially arranged array of microtubules) and associated motor proteins (left panel) vs asters formed in a mitotic extract under the same conditions (right panel, immunofluorescence staining is for tubulin, showing microtubules)
BOTTOM RIGHT: Comparison of an output of the simulation showing conditions under which microtubules are not organized (left panel) vs no organization under the same conditions in a mitotic extract (right panel)